16S/18S/ITS Amplicon Metagenomic Sequencing
- Targeting and identifying the organisms of interest
- Annotating diverse microorganisms in one assay, to obtain a complete composition of microbial communities.
- Detection of pathogens, microorganism contamination, soil bacterial benefits and etc.
OVERVIEW
16S/18S/ITS Amplicon Metagenomic Sequencing is an ultra-deep DNA sequencing method that focuses on sequencing specific target regions (amplicons). Short (<500bp) hypervariable regions of conserved genes or intergenic regions are amplified by PCR and analyzed by next generation sequencing (NGS) technology, to identify and differentiate multiple microbial species from complicated samples. Amplicon metagenomic sequencing is designated to sequence the target genes of 16S ribosomal RNA (rRNA), or 18S rRNA and Internal Transcribed Spacer (ITS) rRNA by universal primers, to describe and compare the phylogeny and taxonomy of bacteria (and archaea) and fungi (such as yeasts, moulds and etc.), respectively.
Our amplicon metagenomic sequencing services provide efficient screen for variants and target organisms, and describe and compare the diversity of multiple complex environments. The approach is frequently used in population and community microbial ecology studies, phylogenetic reconstruction of target microbial groups, identification of individual species in pure cultures, and detection of organisms of interest (pathogens or beneficial), among many others.
What’s Included
- Sample Receipt and Initial QC
- Library Construction and QC
- Sequencing and Data Delivery
- Bioinformatics Analysis
INPUT REQUIREMENTS
- ≥ 200 ng genomic DNA (≥ 10 ng/μL)
- ≥ 1 µg PCR Fragment (≥ 10ng/µL)
Data Deliverable
- OTUs cluster and phylogenetic construction
- Species annotation
- Alpha & Beta diversity analysis
- Comparative analysis
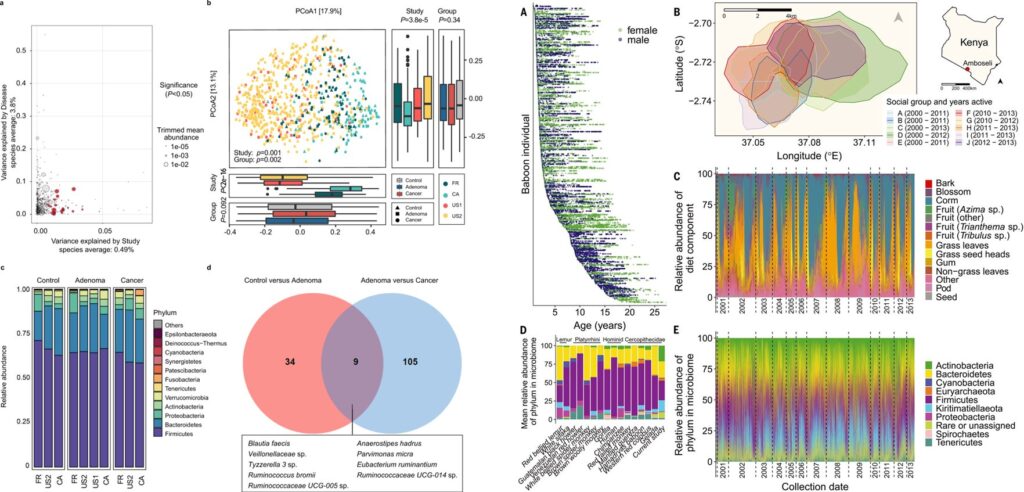
Identification of microbial markers across populations in early detection of colorectal cancer. Nat Commun 2021, 12(1):3063.
Gut microbiome heritability is nearly universal but environmentally contingent. Science 2021, 373(6551):181-186.

